The N3C brings together 16.4B clinical records from 14.3M patients at 74 institutions. To our knowledge, the N3C is the largest national, publicly available HIPAA-limited dataset in US history. The innovative data governance and engineering strategy and public-private-government partnership has made broad sensitive clinical data sharing possible. Clinical informatics has been siloed and competitive; N3C has galvanized sharing of data, methods, and artifacts; furnishing full provenance for rigor, reproducibility, and transparency; and attribution for all types of contributors. The unprecedented availability of this data has catalyzed 376 collaborations collectively involving 3,800+ researchers from 300+ institutions in 25 countries.
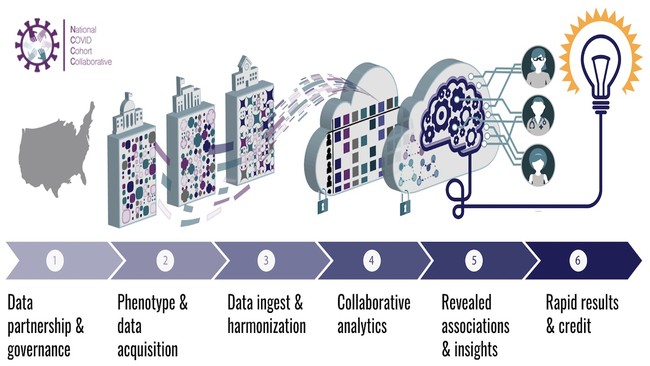
Members diverse in expertise, gender, race, ethnicity, career level, and affiliations are addressing COVID-19 questions in a secure, national Enclave of Electronic Health Record data. We created “workstreams”: A) Data Partnership and Governance (regulatory, data use, code of conduct, attribution policies); B) Phenotype and Data acquisition (inclusion/exclusion criteria and data hand off); C) Data Ingestion and Harmonization (>5000 provenanced data transforms); D) Collaborative Analytics (>30 domain teams, templates, codesets, and training portfolio); and E) Synthetic Data (generation and validation for broader sharing). Multidisciplinary teams (e.g. social determinants of health, kidney, diabetes, machine learning) are supported by training, analytical templates by “logic liaisons”, and codesets and computable phenotypes by “data liaisons.” All contributions are tracked, incentivizing collaboration over competition. Oversight is a first-of-its-kind partnership between NIH and the community. Institutions sign Data Transfer and Use Agreements, individuals obtain IRBs, and NIH provisions access. Governance is created in community meetings; NIH and N3C leadership sign-off to ensure adherence to ethical and regulatory requirements.
To address the lack of centralized patient-level data in the US, the N3C was launched in April 2020 to create a harmonized EHR dataset to identify COVID-19 risk factors, therapies that exacerbate or ameliorate symptoms, and mechanisms. To achieve this urgent and laudable goal, the N3C community came together to create novel governance, transparency, and accountability for clinical data-focused research.
In many data harmonization initiatives, “black box” mapping lacks rules, validation, provenance, or versioning. We scaled manual curation of over 2000 mappings into an automated fully provenanced EHR data harmonization process that is monitorable and correctable as errors or changes to artifacts or data are discovered. Weekly data loads resulted in rapid identification and iteration on data problems and increased knowledge transfer between institutions and the Phenotype and Data acquisition workstream. We implemented a large suite of data quality checks and recovery methods. Each site benefited from seeing its data in the context of the whole, which had never before been realized in the context of distributed research networks. By working together with many sites, we utilized and advanced standards such as OMOP and employ best practices such as a terminology management lifecycle.
We recommend OMOP as the most performant Common Data Model (CDM) for EHR data and encourage the use of source terminologies. We require ORCID for attributing artifacts and manuscripts and use Github and Zenodo/MedRxiv for code, artifact, and document versioning and dissemination. We recommend combining observational data with iterative trial designs and the expanded use of N3C for public health surveillance and collaboration across agencies (NIH, FDA, CDC, DARPA, ARPA-H, HHSS, VA, etc.).
The sheer speed, data volume, scientific productivity, volunteerism, and unique public-private-government partnership is a testament to the N3C community. Models that truly aid clinical research (such as being able to identify long-covid patients) and care (such as changing HIV guidelines) have been realized, as has the first integration of basic research data alongside enough clinical data to enable patient clustering, mechanism identification, and therefore the possibility of greatly enhanced clinical research. The magnitude of data sharing, collaboration, and reuse of code/artifacts/data is unprecedented, and the N3C has even been consulted by the White House for analyses on new therapeutics.
In partnership with distributed EHR research networks, we harmonized four CDMs (OMOP, PCORnet, TriNetX, ACT) and their terminologies (e.g. LOINC, ICD, RXnorm, and SNOMED-CT) and developed a fully provenanced and reproducible pipeline for ingesting, harmonizing, and quality assuring data.
N3C data quality (DQ) review involves automated and manual procedures, internal verification, external validation, conformance, completeness, and plausibility. Beyond well-recognized DQ issues, we discovered heuristics relating to each CDM conformance, demographics, COVID tests, conditions, encounters, measurements, observations, coding completeness, and fitness for use. 66% of sites demonstrated issues and improved their data quality after feedback. We recovered and repaired data that would have been lost to analyses or otherwise provided a poor foundation for analysis in a distributed context. For example, measurements such as mass concentration (mass/volume) are part of a LOINC code but units are not specified. We used the Unified Code for Units of Measure and algorithmic rules to harmonize 88.1% of measurement data with values, and inferred units for 78.2% of measurements with missing units.
Scaling expert curation and mapping yields substantial dividends in terms of the size, quality, and coverage of the resulting dataset. All transformation and analysis code developed by the domain teams are fully reproducible, and artifacts such concept sets are in GitHub for replication and reuse.
While N3C was created to address a public health crisis, in the process we overcame significant barriers to sensitive data sharing and large collaborative science. The N3C created greater rigor, reproducibility, transparency, and better data quality where all organizations benefit. The provenance of all data and analyses has afforded an incredible opportunity to track contributions and transform the previously very competitive and siloed clinical informatics community into a collaborative one, which is illustrated by journals having to index ~200 authors on a manuscript! However, it isn’t only the data sharing, it is the knowledge sharing and reuse that comes along for the ride. Data sharing at this scale enables easier, faster, and more rigorous building on each other's work than traditional manuscript-based dissemination contexts. Conversely, it also enables training opportunities and cross-disciplinary learning that were uncommon before, bringing together clinicians, informaticians, terminologists, epidemiologists, statisticians, computer scientists, basic researchers, patients, and citizen scientists. Such data sharing also helps advance technology and commercial partnerships faster and in a more fit-for-purpose manner.