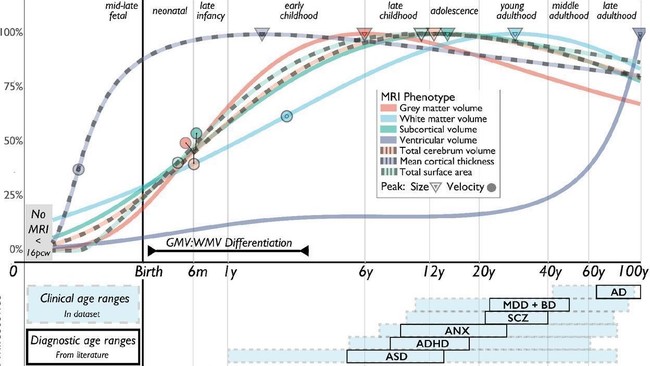
We aimed to create normative brain charts, analogous to pediatric growth charts, for benchmarking individual differences in commonly used neuroimaging metrics over the lifecourse. We coordinated a global consortium to reuse 123,984 MRI scans, from 100+ primary studies, from before birth to 100 years of age. We built and validated statistical models for data harmonization, estimation of normative non-linear growth curves, and individual centile scoring. We discovered and quantified milestones of brain development and abnormal brain aging in Alzheimer’s disease. All methods and results have been openly published with peer review; all tools are openly available via an interactive portal (www.brainchart.io).
Our team grew out of a grass-roots initiative at CHOP/Penn and Cambridge in 2020, motivated to build new research standards for human neuroimaging. The core team aggregated readily available MRI data, then reached out to various research groups to engage them in a global collaborative network. Enabled by Zoom, Slack and Google Drive this network soon grew to include 200+ scientists and to connect many pre-existing consortia. Drs. Seidlitz and Bethlehem worked closely with their institutional legal offices to manage the data sharing effort, and data was stored and processed on the high-performance computing infrastructure at Cambridge. Statistical routines were developed with Dr. Simon White, and the effort was jointly supervised by two physician-scientists, Drs. Aaron Alexander-Bloch and Ed Bullmore. The resulting research tool is a containerised module that allows users to interact with anonymised and group level data in compliance with data access protocols. Users can also benchmark their own datasets anonymously while maintaining privacy of individual-level patient data. This process maximises the utility of the brain charts for the global community while allowing us to iteratively update them with new data as it is shared.
The project started in 2020 and reached the key milestone of open publication in 2022. Our goal was to generate comprehensive life-spanning reference charts of commonly used brain MRI metrics, that could be used to 1) chart normative brain development and aging, and 2) quantify individual (a)typicality of brain structure in terms of centile scores benchmarked by age- and sex-matched norms. In the longer term, we see further development of this technology as crucial for widespread use of standardised quantitative tools (digital neuroradiology) to extract additional information from brain MRI scans to complement the qualitative impressions of neuroradiologists.
It was immediately clear that we needed a very large dataset to accomplish this feat. The project started coincidentally with the onset of the COVID pandemic, which stimulated awareness of the importance of open data and global collaboration. Building on our prior experience with data-sharing initiatives and community-driven science, we adopted an open, transparent and inclusive approach.
We contacted potential collaborators and explained our goals, how their contributions would be appropriately recognised by co-authorship and/or citation of primary datasets, and how our process would comply with any legal or other restrictions that might apply to reuse of their data. Prompt clear communication with collaborators and legal offices at Cambridge and CHOP/Penn was fundamental to successfully aggregating the largest extant collection of human MRI scans. For statistical modeling, we adopted and adapted the WHO-recommended standard for growth curve analysis (GAMLSS), and made all our code and tools openly available, so collaborators could be confident about how their data were analyzed.
We recommend the following practices: 1) an important and timely objective that will be of global value and can only be achieved by data sharing; 2) a clear set of principles concerning appropriate acknowledgement and incentivization of data sharing; 3) diligent attention to the various terms and conditions of data sharing agreements; and 4) total transparency of the scientific process so all collaborators can contribute optimally to the final product.
It is compelling that collectively we were able to rapidly achieve a landmark study of neurodevelopment and aging, which none of us could have delivered by working competitively or independently, with 200+ co-authors harmoniously engaged from start to finish.
We accepted data in any form collaborators were able to share it, converted it to a community-standard data structure for reproducible neuroimaging (the Brain Imaging Data Structure [BIDS]), and processed it with pipelines based on the FreeSurfer library. When encountering restrictive data access, we implemented a federated approach whereby collaborators implemented similar pipelines on their own data. When possible, we obtained permission to openly share datasets for future use by the community. We focused extensively on replicability by creating a technically novel out-of-sample estimator to 1) confirm unbiased and reliable centile scores for scans that had not been included in the reference data, and 2) allow future users to upload their own data for brain charting without any requirement for data transfer agreements or any risk of breaching data confidentiality compliance. For sharing our tools and results, we built an interactive, containerised R Shiny platform (www.brainchart.io) and the fundamental code was openly shared on GitHub (https://github.com/brainchart/Lifespan) for others to use or improve. Our brain chart results can be replicated by other groups applying different approaches to the same underlying data, and new data can be used to validate growth charts using the reproducible software we developed for the original dataset. Looking forward, our brain chart resource can be used as a blueprint for other radiomics applications and other clinical ‘omics data.
Our experience has been overwhelmingly positive. We found a strong appetite across the global MRI research community to share data and we demonstrated convincingly that multiple primary datasets could be harmonized and analyzed to produce an outcome greater than the sum of its parts.
We learnt that even the largest extant MRI dataset is not yet fully representative of socioeconomic and geographic diversity. The majority of available scans were from US or European populations, and generally biased towards ethnically white participants from UMICs. Data sharing on this scale opened our eyes to the need for more inclusive participation in brain research and our tools were designed to allow future studies of currently under-represented demographics to be incorporated in the brain chart framework.
We also learnt that incentives need careful consideration. Some of the most open primary datasets were shared without the relevant investigators satisfying journal criteria for co-authorship, so we had to acknowledge their contributions in other ways. All data-sharing communities should be vigilant about the potential paradox that the most open practices may be least rewarded by traditional standards for significant scientific contribution.